PRODUCTS
With proprietary intellectual property of MicroStacker™ and SuperISH™,
we have a market-leading portfolio of IHC and ISH reagents and instruments.
-
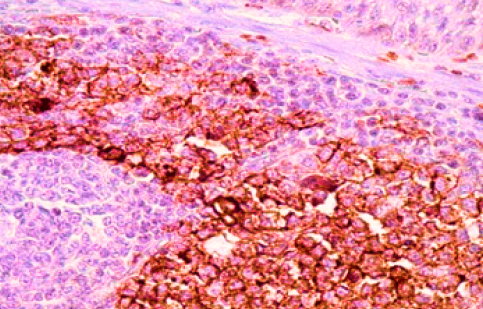 I
IIMMUNOHISTOCHEMISTRY
-
I
FROZEN SECTION IHC
-
I
IMMUNOFLUPRESCENCE
-
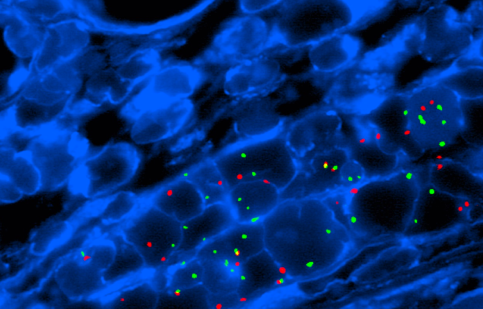 I
IMOLECULAR
-
 I
IINSTRUMENT
-
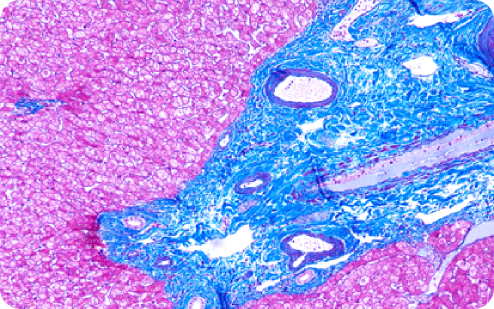 I
IHISTOLOGY
-
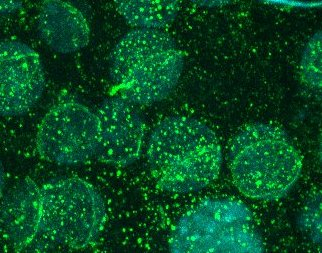 I
IIMMUNOCYTOCHEMISTRY
-
 I
IDIGITAL PATHOLOGY
SOLUTIONS
We provide Clinical Solutions, R&D Collaboration and OEM/ODM Partnership.
FACILITIES & CERTIFICATES
Our facilities in China are NMPA & GMP compliant
and certified for ISO13485 and ISO9001.















 0
0
